4/13 10:00_花田耕介教授_Lineage-Specifically Emerged Genes in Plant Evolution
(Evolutionary Genetics and Genomics)
時間:2023. 04. 13 Thu. 10:00
地 點:跨領域科技研究大樓1樓演講廳
演講者:花田耕介教授
Department of Bioscience and Bioinformatics, Kyushu Institute of Technology, Japan
講題:Lineage-Specifically Emerged Genes in Plant Evolution
主持人:李奇鴻特聘研究員
摘要
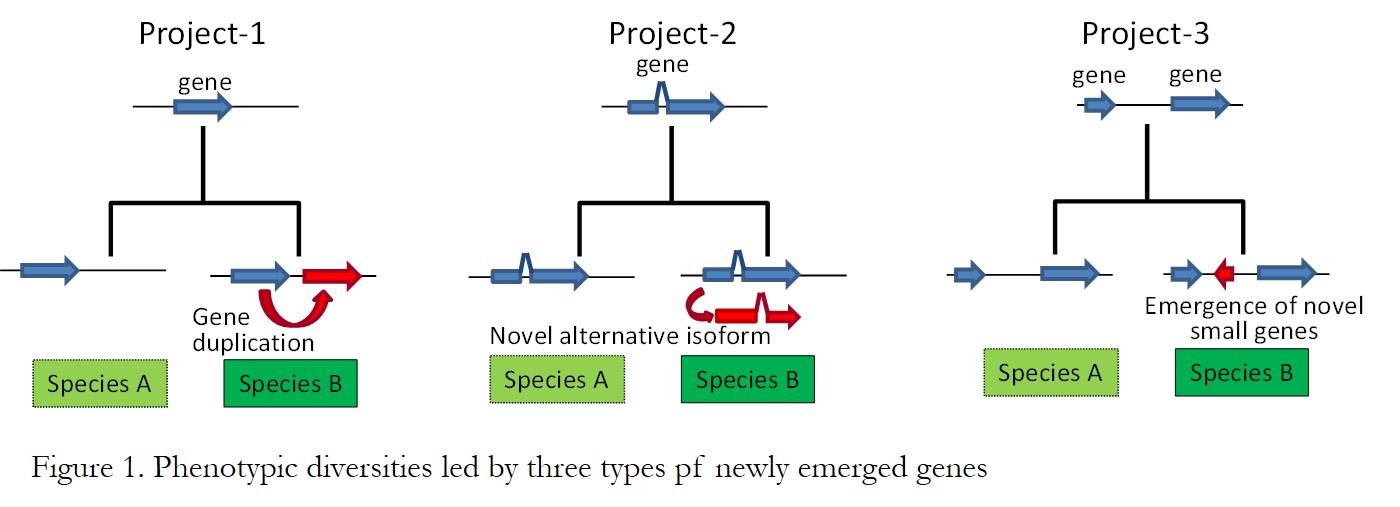
During the course of plant evolution, genomes acquire novel genetic elements as sources of functional diversities. Recent advance of next generation sequencing revealed much of lineage-specifically emerged genes such as duplicated genes, de-novo alternative splicing isoforms and de-novo small coding genes in plant genomes. However, lineage-specifically duplicated genes might keep redundant functions with original ones. Also, de-novo alternative splicing isoforms and small genes are expressed as RNA, but the RNA products might be just transcription errors. Therefore, I’m trying to identify lineage-specific genes inducing phenotypic diversities in plants. I mainly used the best model organism of plants (Arabidopsis thaliana) to understand the adaptive evolution throughout lineage-specifically emerged genes. First, I collected various omics data such as transcriptome, proteome, metabolome, interactome, phenome, etc in A. thaliana (Evaluation). Throughout the integration of these omics by bioinformatics analysis, I tried to infer recently emerged genes causing phenotypic changes in evolution (Study & Design). Then, I examined the functional roles of inferred genes by transgenic plants (Construction). This strategy (Evaluation→Study→Design→Construction) is useful to understand the molecular mechanisms of target genes. Here, as sources of phenotypic diversities, I will introduce the evolution of phenotypic variation by three types of newly emerged genes such as gene duplication, alternative splicing isoforms, and small genes (Figure 1).
~歡迎踴躍參加~
~與會者建議配戴口罩~




